If something is amiss, you know what has gone wrong rather than trying “hundreds of ways. Visually verify the correctness of the coordinates after every step. Proceed as normal with positioning the complex within a box, solvate, etc. Update the number of atoms in the second line of o. > example autodock vina output is attached (its a conformer of the ace > native ligand dude). 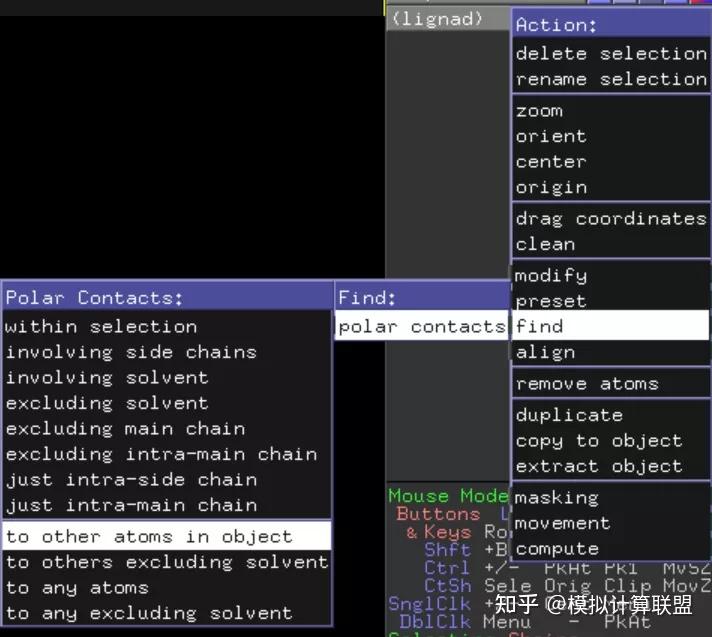
Strictly speaking, this isn’t necessary (you can use PDB format) but if any H were built on to the protein via pdb2gmx this may be easier:Ĭopy and paste the coordinates of lig.gro into o and save as o. > the autodock pdbqt file has only the polar hydrogens in it part of the > trick is to re-add the hydrogens. Change coordinate format for the ligand.Generate ligand topology with whatever tools are available for the force field you’ve chosen to use. Here, we converted the ligands files into PDBQT file format using PyRX and Open Bable software, which is freely available online. Save the ligand coordinates (assumes it has all the H atoms you need, if not, you need to build those on in some modeling software and ensure it does not alter the non-H coordinates):.Separate the protein and ligand from the original docked complex (assumes “LIG” is the ligand name, just substitute whatever it really is):.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
